SARS-CoV-2
Diaclone continues to innovate and provide you with new research tools...
Diaclone proposes a range of recombinant proteins and monoclonal antibodies specific to SARS-CoV-2 to help you develop your own serological assay or other projects against Covid-19.
RECOMBINANT HUMAN SARS-CoV-2 PROTEINS
Recombinant proteins for your research against SARS-CoV-2 and other coronaviruses: Spike protein, Nucleoprotein,…. and other variants under development.
|
Specificity |
Expression system |
Tag |
Size (100µg) |
Other sizes |
|
Recombinant Human SARS-CoV-2 Nucleoprotein Full length (1-419) |
HEK293 cells |
HIS Tag C-Terminus |
* |
|
|
Recombinant Human SARS-CoV-2 Nucleoprotein Full length (1-419) |
E. coli |
HIS Tag C-Terminus |
* |
|
|
Recombinant Human SARS-CoV-2 Spike S1 RBD (Active) |
HEK293 cells |
HIS Tag C-Terminus |
* |
|
|
Recombinant Human Angiotensin-converting enzyme 2 (ACE2) Extracellular Domain (19-615) |
HEK293 cells |
HIS Tag C-Terminus |
* |
|
|
HEK293 cells |
HIS Tag C-Terminus |
* | ||
|
HEK293 cells |
HIS Tag C-Terminus |
* | ||
|
HEK293 cells |
HIS Tag C-Terminus |
* | ||
|
HEK293 cells |
HIS Tag C-Terminus |
* | ||
|
HEK293 cells |
HIS Tag C-Terminus |
* | ||
|
HEK293 cells |
HIS Tag C-Terminus |
* | ||
|
HEK293 cells |
Human IgG1 Fc |
* |
|
 |
ANTI-SARS-CoV-2 MONOCLONAL ANTIBODIES
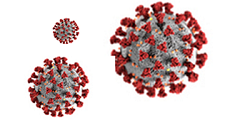
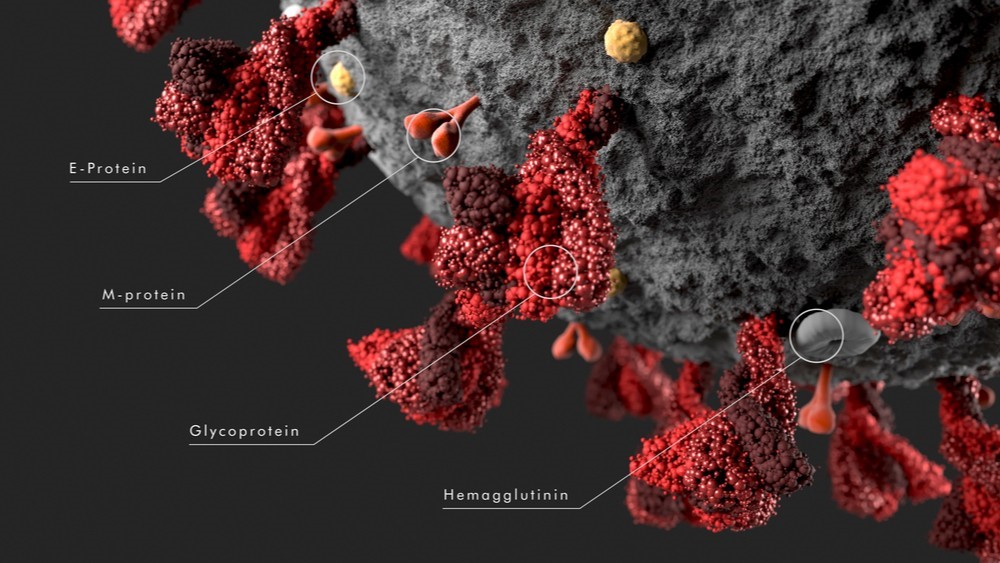
Coronaviruses are positive-sense RNA viruses having an extensive range of natural hosts and typically affect the respiratory system. SARS-CoV-2 (Severe acute respiratory syndrome coronavirus 2) has four structural proteins, known as the S (spike), E (envelope), M (membrane), and N (nucleocapsid) proteins; the N protein holds the RNA genome, and the S, E, and M proteins together create the viral envelope.
The S protein is a large viral transmembrane protein associated in a trimer manner on the virion surface, giving the virion a corona or crown-like appearance. Functionally it is the protein responsible for allowing the virus to attach to and fuse with the membrane of a host cell via the ACE2 protein. The ectodomains in all CoVs S proteins have similar domain organizations, divided into two subunits: the first one, S1, helps in host receptor binding, while the second one, S2, accounts for fusion. It also acts as a critical factor for tissue tropism and the determination of host range. The S protein is one of the proteins of CoVs capable of inducing most important host immune responses.
Standard sizes and Bulk formats available
|
Description |
Target species |
Clone |
Synonym |
Read more |
|---|---|---|---|---|
|
Anti-SARS-CoV-2 RBD domain Azide Free |
SARS-CoV-2 |
B-B60 |
Receptor binding domain Spike protein RBD S1 protein |
|
|
Anti-SARS-CoV-2 RBD domain Azide Free |
SARS-CoV-2 |
B-D69 |
Receptor binding domain Spike protein RBD S1 protein |
|
|
Anti-SARS-CoV-2 RBD domain Azide Free |
SARS-CoV-2 |
B-R41 |
Receptor binding domain Spike protein RBD S1 protein |
|
| Anti-SARS-CoV-2 RBD domain Azide Free | SARS-CoV-2 | B-K45 |
Receptor binding domain Spike protein RBD S1 protein |
Detail |
| Anti-SARS-CoV-2 RBD domain Azide Free | SARS-CoV-2 | B-A67 |
Receptor binding domain Spike protein RBD S1 protein |
Detail |
RECOMBINANT ANTI-SARS-CoV-2 ANTIBODIES
Over the years, Diaclone has invested in recombinant antibody technologies to complete our range of monoclonal antibodies. Alongside our monoclonal antibody development platform by conventional hybridoma technology, Diaclone is now able to develop recombinant antibodies by optimized Phage Display technology.
Standard sizes and Bulk formats available
|
Description |
Target species |
Clone |
Synonym |
Read more |
|---|---|---|---|---|
|
Anti-SARS-CoV-2 Nucleoprotein Azide Free |
SARS-CoV-2 |
CR3018 |
Nucleoprotein, SARS-CoV nucleoprotein, Nucleocapsid protein |
|
|
Anti-SARS-CoV-2 Spike S1 Protein Azide Free |
SARS-CoV-2 |
CR3022-IgG |
Receptor binding domain Spike protein, RBD spike protein |
|
|
Anti-SARS-CoV-2 Spike S1 Protein Azide Free (IgA) |
SARS-CoV-2 |
CR3022-IgA |
Receptor binding domain Spike protein, RBD spike protein |
info [at] diaclone [dot] com ( Contact us to test these new products, Subject: SARS-CoV-2 specific mAbs, ) or simply for more information
Documentation
Ideal for your Covid-19 Studies
Development of SARS-CoV-2 RDB Protein variants
Diaclone - Editos
Complete Proteome of COVID-19
The genome of SARS-CoV-2 counts 29,811 nucleotides, encoding for 29 different proteins. The translation of the linear single-stranded RNA conducts to the generation of the following proteome:
- 4 are structural proteins, S, N, M, and E
- 16 proteins are non-structural proteins or NSP: the first 11 are encoded in ORF1a whereas the last 5 are encoded in ORF1b
- 9 are accessory proteins named ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8, ORF9b, ORF9c, and ORF10
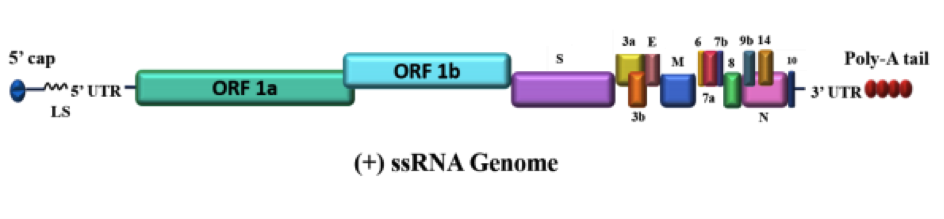
Figure 1. Genome architecture of SARS-CoV-2. SARS-CoV-2 contains a positive single-stranded mRNA as genetic material. The 5’ capped mRNA has a leader sequence (LS), poly-A tail at 3’ end, and 5’ and 3’ UTR. It contains the following genes: ORF1a, ORF1b, Spike (S), ORF3a, ORF3b, Envelope (E), Membrane (M), ORF6, ORF7a, ORF7b, ORF8, ORF9b, ORF14, Nucleocapsid (N), and ORF10. Dark Proteome of Newly Emerged SARS-CoV-2 in Comparison with Human and Bat Coronaviruses. Rajanish Giri, Taniya Bhardwaj, Meenakshi Shegane, Bhuvaneshwari R. Gehi, Prateek Kumar, Kundlik Gadhave.
1. Structural proteins of SARS-COV-2
- Spike (S). Trimeric transmembrane spike glycoprotein (1,273 a.a.) precursor is cleaved into S1 and S2 segments in order to bind to ACE2 receptors. A mutation in the SARS-CoV-2 S protein allows the human enzyme called furin to do the proteolytic cleavage into the sequence (PRRARS|V). Since bat CoV doesn’t present this sequence, its apparition may be an explanation for the virus ability to infect humans (1).
- Nucleocapsid (N). Dimer (419 a.a.) that binds viral genome to membrane forming a shell and demonstrates non-specific nucleic acid binding capability (2). Although the overall structure is similar to other reported coronavirus nucleocapsids, the surface electrostatic potential characteristics are unique (3).
- Membrane (M). Matrix protein (222 a.a.) is the most abundant structural component of the virion and defines the shape of the envelope. M protein of SARS-CoV-2 has a triple helix bundle, forms a single 3-transmembrane domain (TM), and is homologous to the prokaryotic sugar transport protein semiSWEET. The advantage and role of sugar transporter-like structures in viruses are still unknown (4). M protein cooperates with Spike during the cell attachment and entry (5).
- Envelope (E). This small membrane protein (75 a.a.) is involved in viral assembly, budding, and pathogenesis. Envelope protein is identical to its counterparts from CoV MP798, CoVZXC21, CoVZC45 and RaTG13. However, a substitution at position 69, and a deletion in position 70 were found (5).
2. Non-structural proteins of SARS-COV-2
Non-Structural Proteins (nsp) are expressed as two long polyproteins (pp1a and pp1ab) then are cleaved into 16 mature smaller proteins by the papain-like protease (PLpro) and the 3-chymotrypsinlike protease (3CLpro, also known as the main protease-Mpro).
- Nsp1 (180 aa): 28 inserts or deletions occurs along its amino-acids sequence, compared to SARS-Cov. The role of nsp1 is believed to be very similar in SARS-Cov and SARS-Cov-2, suppressing type I IFN expression (6).
- Nsp2 (638 aa): nsp 2 interacts with a host protein complex of PHB1 and PHB2 involved in mitochondrial biogenesis (7). Position 321 of this methyltransferase sequence has a polar amino acid while Bat SARS‐like coronavirus has a non-polar amino acid. It can be speculated, that due to this polarity, and potential to form H‐bonds, the glutamine amino acid may confer higher stability to the protein (8).
- Nsp3 (1,945 aa): Nsp3 shuts down host enzymes called PARPs, which prevent viruses from replicating (9). The destabilizing mutation in nsp3 proteins could suggest a potential mechanism differentiating COVID‐2019 from SARS (8).
- Nsp4 (500 aa): Nsp4 critically interacts with nsp3 to rearrange host cell membrane. Only their synergy is able to perform the job (10).
- Nsp5 (306 aa): the main protease of 306 aa cleaves at 11 sites. COVID-19 NSP5 is also highly homologous to SARS NSP5 (96% identity, 98% similarity) (11).
- Nsp6 (290 aa): It interacts with nsp3 and nsp4. Nsp6 presents 7 putative trans-membrane helices like in other coronaviruses. The presence of two mutations affecting the Non-Structural Protein 6 has been recently found (12).
- Nsp7 (83 aa) & Nsp 8 (198 aa): The SARS-coronavirus nsp7+nsp8 primase complex is capable of both de novo initiation and primer extension (13). The complex triggers RNA-synthesizing activity of nsp12 (14). NS7b and NS8, were exclusively conserved among 2019-nCoV, BetaCoV_RaTG, and BatSARS-like Cov. Functional changes in the NS7b and NS8 proteins during evolution may provide important information to explore the human infective property of 2019-nCoV (15).
- Nsp9 (113 aa): Current understanding suggests that Nsp9 is involved in viral genomic RNA reproduction. The structure of the SARS-CoV-2 Nsp9 revealed the high level of structural conservation within the Nsp9 family (16).
- Nsp10 (139 aa): forms a complex with NSP16 to cap viral mRNA transcripts for efficient translation and to evade immune surveillance (17).
- Nsp11 (13 aa): overlapping sequence with nsp10, nsp11 short peptide function is still unknown.
- Nsp12 (932 aa): RNA polymerase complex consisting of the viral RdRp (nsp12) and associated cofactors (nsp7 and nsp8). Alignment of nsp12 for the whole CoV family indicates that the SARS-CoV-2 nsp12 is almost identical to that of the SARS-CoV (96% identity, 98% similarity) (18).
- Nsp13 (601 aa): Like SARS and MERS-Nsp13, the overall structure of SARS-CoV-2 nsp13 adopted a triangular pyramid shape comprising five domains (19). Nsp13 is a helicase that unpacks viral genome material to make it more accessible.
- Nsp14 (527 aa): Nsp14 is the proofreading non-structural protein for normal CoV recombination. Thus, nsp14-ExoN is a key determinant of both high fidelity CoV replication and recombination, and thereby represents a highly-conserved and vulnerable target for virus inhibition and attenuation (20).
- Nsp15 (346 aa): nidoviral RNA uridylate‐specific endoribonuclease (NendoU). While initially Nsp15 was thought to directly participate in viral replication, it was later shown that Nsp15‐deficient coronaviruses were viable and replicating. More recently, it was proposed that NendoU activity of Nsp15 is responsible for the protein interference with the innate immune response, though other studies indicate that the process is independent of the endonuclease activity. There are also suggestions that Nsp15 degrades viral RNA to hide it from the host defences (21).
- Nsp16 (298 aa): forms a complex with NSP10 to cap viral mRNA transcripts for efficient translation and to evade immune surveillance (17).
3. Accessory proteins of SARS-COV-2
- ORF3a (275 aa): the 3a protein is unique to SARS-CoV and is essential for virulence, infectivity, ion channel formation, and virus release (22).
- ORF3b (22 aa): overlapping sequence with 3a, the 3b short peptide is a potent IFN-1 antagonist but allegations have yet to be confirmed. The ORF3b sequences of SARS-CoV-2 are considerably shorter than those of their SARS-CoV orthologs (153.2 ± 0.47 amino acids on average) (23).
- ORF6 (61 aa): IFN-1 antagonist. ORF6 has been shown to be an IFN antagonist that disrupts karyopherin transportation of transcriptions factors like STAT1 (24, 25).
- ORF7a (121 aa): SARS-CoV ORF7a directly binds to BST-2 and inhibits its activity by blocking the glycosylation of BST-2 (26).
- ORF7b (43 aa): overlapping sequence with 7a, ORF7b protein is not only an accessory protein but a structural component of the SARS-CoV virion (27). We haven’t found any reference in SARS-CoV-2 so far.
- ORF8 (121 aa): ORF8 is the most different protein compared to SARS-Cov (30% homology). A 382-nt deletion covering the ORF8 of SARS-CoV-2 has been reported. The deletion also removes the ORF8 transcription-regulatory sequence (TRS), which in turn enhances the downstream transcription of the N gene. (28)
- ORF9b (97 aa): coded within the N gene, ORF9b interacts with a mitochondrial import receptor, Tom70, which acts as an essential adaptor linking MAVS to TBK1/IRF3, resulting in the activation of IRF-3 (29).
- ORF9c (XX aa): coded within the N gene, sigma 2 receptors are hijacked by ORF9c. ORF9c protein was found to interact with multiple proteins that modulate IkB kinase and NF-kB signalling pathway including NLRX1, F2RL1, NDFIP2 (29).
- ORF10 (38 aa): ORF10 does not have any similar proteins in the NCBI repository for SARS-CoV (ORF10 had a premature stop codon in both SARSCoV and BM48-31) and seems unique to SARS-CoV-2 (30).
And you, which one of these proteins are you working on?
Which one would you like to see developed and validated?
You can now contact Diaclone to discuss these topics!
REFERENCES
(1) COVID-19 Coronavirus spike protein analysis for synthetic vaccines, a peptidomimetic antagonist, and therapeutic drugs, and analysis of a proposed achilles’ heel conserved region to minimize probability of escape mutations and drug resistance. B. Robson
(2) Biochemical characterization of SARS-CoV-2 nucleocapsid protein Weihong Zeng, Guangfeng Liu, Huan Ma, Dan Zhao, Yunru Yang, Muziying Liu, Ahmed Mohammed, Changcheng Zhao, Yun Yang, Jiajia Xie, Chengchao Ding, Xiaoling Ma, Jianping Weng, Yong Gao, Hongliang He, and Tengchuan Jina
(3) Crystal structure of SARS-CoV-2 nucleocapsid protein RNA binding domain reveals potential unique drug targeting sites. Acta Pharmaceutica Sinica B. 1016/j.apsb.2020.04.009.Kang, Sisi & Yang, Mei & Hong, Zhongsi & Zhang, Liping & Huang, Zhaoxia & Chen, Xiaoxue & He, Suhua & Zhou, Ziliang & Zhou, Zhechong & Chen, Qiuyue & Yan, Yan & Zhang, Changsheng & Shan, Hong & Chen, Shoudeng.
(4) Thomas, S. The Structure of the Membrane Protein of SARS-CoV-2 Resembles the Sugar Transporter semiSWEET
(5) Bianchi, Martina & Benvenuto, Domenico & Giovanetti, Marta & Angeletti, Silvia & Ciccozzi, Massimo & Pascarella, Stefano. (2020). Sars-CoV-2 Envelope and Membrane Proteins: Differences from Closely Related Proteins Linked to Cross-species Transmission?
(6) Severe Acute Respiratory Syndrome Coronavirus nsp1 Suppresses Host Gene Expression, Including That of Type I Interferon, in Infected Cells. Krishna Narayanan, Cheng Huang, Kumari Lokugamage, Wataru Kamitani, Tetsuro Ikegami, Chien-Te K. Tseng, Shinji Makino
(7) Comparative Genomic Analysis of Rapidly Evolving SARS CoV-2 Viruses Reveal Mosaic Pattern of Phylogeographical Distribution Roshan Kumar, Helianthous Verma, Nirjara Singhvi , Utkarsh Sood , Vipin Gupta, Mona Singh , Rashmi Kumari, Princy Hira, Shekhar Nagar, Chandni Talwar, Namita Nayyar, Shailly Anand, Charu Dogra Rawat, Mansi Verma, Ram Krishan Negi, Yogendra Singh and Rup Lal.
(8) COVID‐2019: The role of the nsp2 and nsp3 in its pathogenesis. Silvia Angeletti, Domenico Benvenuto, Martina Bianchi, Marta Giovanetti, Stefano Pascarella, Massimo Ciccozzi
(9) https://timesofindia.indiatimes.com/india/a-covid-guide-understanding-the-s-factor/articleshow/75671621.cms
(10) Two-amino acids change in the nsp4 of SARS coronavirus abolishes viral replication. Yusuke Sakaiad, Kengo Kawachiab, Yutaka Teradaa, Hiroko Omori, Yoshiharu Matsuur, Wataru Kamitani
(11) Global profiling of SARS-CoV-2 specific IgG/ IgM responses of convalescents using a proteome microarray He-wei Jiang, Yang Li, Hai-nan Zhang, Wei Wang, Dong Men, Xiao Yang, Huan Qi, Jie Zhou, Sheng-ce Tao
(12) Evolutionary analysis of SARS-CoV-2: how mutation of Non-Structural Protein 6 (NSP6) could affect viral autophagy. Domenico Benvenuto, Silvia Angeletti, Marta Giovanetti, Roberto Cauda, Massimo Ciccozzi, Antonio Cassone.
(13) The SARS-coronavirus nsp7+nsp8 complex is a unique multimeric RNA polymerase capable of both de novo initiation and primer extension. Aartjan J.W. te Velthuis, Sjoerd H. E. van den Worm, and Eric J. Snijder.
(14) One severe acute respiratory syndrome coronavirusprotein complex integrates processive RNApolymerase and exonuclease activitiesLorenzo Subissia, Clara C. Posthumab, Axelle Colleta, Jessika C. Zevenhoven-Dobbeb, Alexander E. Gorbaleny, Etienne Decroly, Eric J. Snijder, Bruno Canarda, and Isabelle Imbert
(15) Nonstructural proteins NS7b and NS8 are likely to be phylogenetically associated with evolution of 2019-nCoV. Fahmi, Muhamad & Kubota, Yukihiko & Ito, Masahiro
(16) Crystal structure of the SARS-CoV-2 non-structural protein 9, Nsp9. D. R. Littler, B. S. Gully, R. N. Colson, J Rossjohn.
(17). Rosas-Lemus, Monica & Minasov, George & Shuvalova, Ludmilla & Inniss, Nicole & Kiryukhina, Olga & Wiersum, Grant & Kim, Youngchang & Jedrzejczak, Robert & Enders, Michael & Jaroszewski, Lukasz & Godzik, Adam & Joachimiak, Andrzej & Satchell, Karla. (2020). The crystal structure of nsp10-nsp16 heterodimer from SARS CoV-2 in complex with S-adenosylmethionine.
(18) Remdesivir and SARS-CoV-2: Structural requirements at both nsp12 RdRp and nsp14 Exonuclease active-sites. AshleighShannona, Nhung Thi-Tuyet Le, Barbara Seliskoa, Cecilia Eydoux, Karine Alvarez, Jean-Claude Guillemot, Etienne Decroly, Olve Peersen, Francois Ferron, BrunoCanard
(19) Structural elucidation of SARS-CoV-2 vital proteins: Computational methods reveal potential drug candidates against main protease, Nsp12 polymerase and Nsp13 helicase. Muhammad Usman Mirza, Matheus Froeyen.
(20) The coronavirus proofreading exoribonuclease mediates extensive viral recombination. Gribble, Jennifer & Pruijssers, Andrea & Agostini, Maria & Anderson-Daniels, Jordan & Chappell, James & Lu, Xiaotao & Stevens, Laura & Routh, Andrew & Denison, Mark.
(21) Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2. Youngchang Kim, Robert Jedrzejczak, Natalia I. Maltseva, Mateusz Wilamowski, Michael Endres, Adam Godzik, Karolina Michalska, Andrzej Joachimiak.
(22) SARS-CoV-2 and ORF3a: Nonsynonymous Mutations, Functional Domains, and Viral Pathogenesis - Elio Issa, Georgi Merhi, Balig Panossian, Tamara Salloum, Sima Tokajian
(23) SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is further increased by a naturally occurring elongation variant. Yoriyuki Konno, Izumi Kimura, Keiya Uriu, Masaya Fukushi, Takashi Irie, Yoshio Koyanagi, So Nakagawa, Kei Sato.
(24) Severe acute respiratory syndrome coronavirus open reading frame (ORF) 3b, ORF 6, and 278 nucleocapsid proteins function as interferon antagonists. Kopecky-Bromberg, S.A., Martinez-Sobrido, L., Frieman, M., Baric, R.A. & Palese.
(25) Frieman, M., et al. Severe acute respiratory syndrome coronavirus ORF6 antagonizes 280 STAT1 function by sequestering nuclear import factors on the rough endoplasmic 281 reticulum/Golgi membrane. Journal of virology 81, 9812-9824 (2007).
(26) Severe acute respiratory syndrome coronavirus ORF7a inhibits bone marrow stromal antigen 2 virion tethering through a novel mechanism of glycosylation interference. J.K. Taylor, C.M. Coleman, S. Postel, J.M. Sisk, J.G. Bernbaum, T. Venkataraman, et al.
(27) The ORF7b protein of severe acute respiratory syndrome coronavirus (SARS-CoV) is expressed in virus-infected cells and incorporated into SARS-CoV particles. Schaecher SR, Mackenzie JM, Pekosz A
(28) Discovery of a 382-nt deletion during the early evolution of SARS-CoV-2. Running title: A SARS-CoV-2 deletion variant. Yvonne CF Su, Danielle E Anderson, Barnaby E Young, Feng Zhu, Martin Linster, Shirin Kalimuddin, Jenny GH Low, Zhuang Yan, Jayanthi Jayakumar, Louisa Sun, Gabriel Z Yan, Ian H Mendenhall, Yee-Sin Leo, David Chien Lye, Lin-Fa Wang, Gavin JD Smith.
(29) A SARS-CoV-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential DrugRepurposing David E. Gordon, Gwendolyn M. Jang, Mehdi Bouhaddou, Jiewei Xu, Kirsten Obernier, Matthew J. O'Meara, Jeffrey Z. Guo.
(30) On the origin and continuing evolution of SARS-CoV-2. Xiaolu Tang, Changcheng Wu, Xiang Li, Yuhe Song, Xinmin Yao, Xinkai Wu, Yuange Duan, Hong Zhang, Yirong Wang, Zhaohui Qian, Jie Cui, and Jian Lu.



